GenePlaza 30 APR, 2018
WORLD PREMIERE AT GENEPLAZA
Where did your Ancestors live 8.000 years ago?
The K14 Ancient Cultures Admixture Calculator will give you the answer!
We are very proud at GenePlaza to have the world’s first DNA based application to include the last genomes of neolithic ancestral lately discovered!
Those are groundbreaking discoveries allowing the developer of this application, Mr Khan, to have a better understanding of our ancestry back to 8.000 years ago.
The ADMIXTURE based calculator parses your genome and compares it to very recently sequenced ancient cultures from Europe, Africa, Central and South Asia.
Mr Khan said about his last K14 application:
“These new higher quality genomes greatly added to our understanding of the population demography of Europe, Western, Central, and South Asia.
Thus we believe that this is the most accurate Ancestry based calculator to date which is based on ancient populations.”
The main difference with the creator’s Ancient ADMIXTURE calculator is that the latter breaks down ethnogenesis in deeper neolithic ancestral terms than this calculator
How to get this study with your own data?
- You don’t have your DNA
- You have your DNA
A quick overview of the K14 Ancient Cultures Admixture Calculator
THE K14 ANCIENT CULTURES CALCULATOR
The motivation behind this calculator was the recent publication of dozens of higher quality ancient genomes in Mathieson et al., 2018; The Genomic History Of Southeastern Europe and in Olalde et al., 2017 The Beaker Phenomenon And The Genomic Transformation Of Northwest Europe, and in Narasimhan et al., 2018,The Genomic Formation of South and Central Asia.
These genomes greatly added to our understanding of the population demography of Europe, Western, Central, and South Asia, and this calculator is the 1st public ADMIXTURE based calculator to parse your genome and compare it to very recently sequenced ancient cultures from Central and South Asia.
To increase the accuracy of the results, and SNP overlaps between the customer and the calculator population references, only genomes with the highest average read depth coverage were carefully chosen to source the component allele frequencies.
The calculator algorithm used is detailed at the calculator creator’s website; Eurasian DNA.
This calculator uses higher coverage ancient genomes from the aforementioned as well as previous studies to represent the various Neolithic, Chalcolithic, and Bronze Age cultures, stretching from western Europe all the way to central Asia and Siberia, and which contributed to the genetic makeup of the various modern populations currently residing across Eurasia.
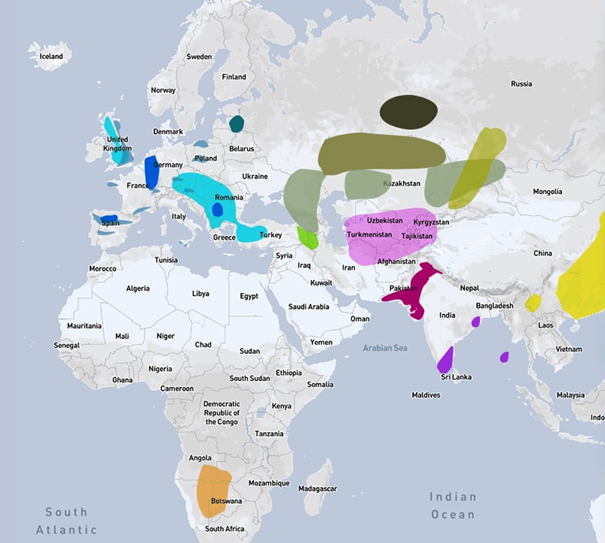
A map of the various ancient cultures existing up to 8000 years ago

Beaker people are known for their distinctive bell beaker style pottery. The culture spread across Europe likely from the Iberian peninsula all the way to Poland around 4700 years ago and lasted till around 3800 years ago. They appear to have displaced the Corded Ware culture which had thrived earlier in eastern Europe.
Prior to the spread of the Beaker culture, Britain was occupied by British Neolithic farmers who were genetically very similar to Iberian Neolithic farmers, suggesting a movement to Britain from Western Europe rather from Germany. The German and SE European Neolithic farmers are likely ancestral to the Iberian and British Neolithic farmers, the latter being distinguished by an additional layer of Western-European Hunter-Gatherers (WHG) admixture. WHG were the long-time occupants of Europe prior to the arrival of the Neolithic farmers from the Near-East around 8000 years ago.
WHG who had occupied Europe for many millennia since the Upper Paleolithic appear to have survived in almost un-admixed form until as recently as 7800 years ago in Serbia and Romania (Iron Gates HG). They were subsequently absorbed into the Neolithic farmer societies which had spread from Anatolia into Europe around 8000 years ago.
The 2nd major population movement into Europe came from the Eurasian steppes (Russia) to the east around the Bronze age. These Eurasian steppe folks were derived from cultures such as the Yamna, Srubna, and Andronovo and it is very likely that is how Indo-European languages were introduced into Europe. Todays Europeans are substantially a tri-fold mixture of WHG, Neolithic farmers originating from the Near-East, and Eurasian steppe pastoralists, in varying proportions. Eurasian steppe pastoralists genetic sub-structure includes Eastern European Hunter-Gatherers (EHG) ancestry as well as Caucasus Hunter-Gatherers (CHG) and Iranian Neolithic farmer ancestry.
There are also additional other minor contributions to Europe’s genetic landscape, including minor African/SW Asian admixture especially in southern Europe, and E Asian / Siberian admixture, which was likely contributed by populations related to the Bronze age Karasuk culture via Uralic proxies, Scythians, and various Turkic groups.
List of the ancient cultures referred to in this app

Here we also utilize for the first time ancient genomes from Turan (present-day Iran, Afghanistan, Uzbekistan, Turkmenistan, and Tajikistan), the Indus Valley ( present-day Pakistan), and from various cultures in Kazakhstan and surrounds, which are believed to have introduced Indo-European languages into the region. Ancestry from the Eurasian Steppe genetically linked Europe and South Asia in the Bronze Age. Present day South and West Asians owe their existence to these ancient populations.
From the Narasimhan et al. 2018 pre-print
Our data reveal a complex set of genetic sources that ultimately combined to form the ancestry of South Asians today. We document a southward spread of genetic ancestry from the Eurasian Steppe, correlating with the archaeologically known expansion of pastoralist sites from the Steppe to Turan in the Middle Bronze Age (2300-1500 BCE). These Steppe communities mixed genetically with peoples of the Bactria Margiana Archaeological Complex (BMAC) whom they encountered in Turan (primarily descendants of earlier agriculturalists of Iran).
Steppe communities integrated farther south throughout the 2nd millennium BCE, and we show that they mixed with a more southern population that we document at multiple sites as outlier individuals exhibiting a distinctive mixture of ancestry related to Iranian agriculturalists and South Asian hunter-gathers. We call this group Indus Periphery because they were found at sites in cultural contact with the Indus Valley Civilization (IVC) and along its northern fringe, and also because they were genetically similar to post-IVC groups in the Swat Valley of Pakistan.
By co-analyzing ancient DNA and genomic data from diverse present-day South Asians, we show that Indus Periphery-related people are the single most important source of ancestry in South Asia—consistent with the idea that the Indus Periphery individuals are providing us with the first direct look at the ancestry of peoples of the IVC—and we develop a model for the formation of present-day South Asians in terms of the temporally and geographically proximate sources of Indus Periphery-Our results show how ancestry from the Steppe genetically linked Europe and South Asia in the Bronze Age, and identifies the populations that almost certainly were responsible for spreading Indo-European languages across much of Eurasia.
Have a look at the preview of the K14 results on GenePlaza and Buy it after creating your account if you have your DNA or buy your DNA kit if you don’t have it yet.
The researcher who developed this K14 Cultures Admixture Calculator is Mr Khan. Visit the website of the author: EurasianDNA.com
How does it work?
This app is dedicated to furthering the understanding of population histories. To facilitate this, the author, Mr Khan, employs an arsenal of tools, including programs such as:
1- ADMIXTOOLS by Reich Lab from David Reich: To formally analyze genomes for shared genetic drift, and for modelling samples based on ancestral components, (qpDstat, qpAdm, f3, etc);
2- PLINK: To prepare and process DNA data sets;
3- BEAGLE: For phasing, imputation and IBD analysis;
4- ADMIXTURE: For grouping individuals into clusters based on shared allele frequencies for some derived alleles.
Do you want to read more?
Have a look at this Guardian article: Arrival of Beaker folk changed Britain forever, ancient DNA study shows
Talking about us:
Two websites for sharing and comparing your results: